How can a country build its own 1000 Genomes Project?
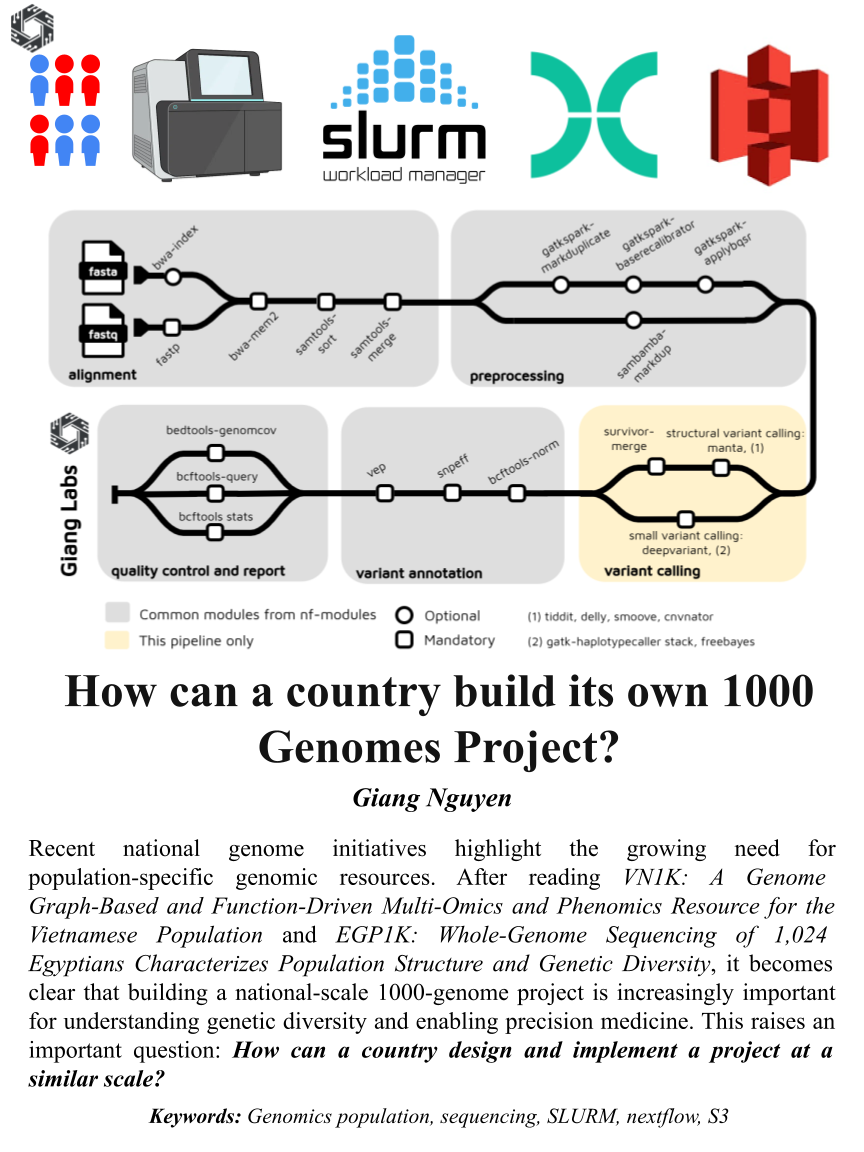
Recent national genome initiatives highlight the growing need for population-specific genomic resources. After reading VN1K: a genome graph-based and function-driven multi-omics and phenomics resource for the Vietnamese population and EGP1K: Whole-Genome Sequencing of 1,024 Egyptians Characterizes Population Structure and Genetic Diversity, it becomes clear that building a national-scale 1000-genome project is increasingly important for understanding genetic diversity, improving disease research, and enabling precision medicine.